I use $30K PK software every day. So I built my own.
A pharmacist's case for why non-commercial pharmacokinetic analysis software is long overdue and what it takes to build one that validates correctly.
More in Build Notes
- The VORA Overhaul: Dropping Real-Time Q&A, Building Human-in-the-Loop Memos, and a Three-Column Layout
- Mobile-First Playground: Making an Astrology Grid Actually Work on a Phone (And Go Viral While Doing It)
- How I Fixed AI Over-correction
- I Spent 3 Hours Trying to Proxy a Blog Subdomain. Here's My Descent Into Madness.
- The i18n + SEO Cleanup Chronicles: Canonical Chaos, hreflang Therapy, and Other Adventures
A colleague in clinical research asked me: "Do you think someone could build a decent NCA tool for under $1,000?"
My first instinct was to rattle off every reason why not. Regulatory compliance. Numerical precision. Data traceability. WinNonlin has a 30-year head start and costs about $30K.
My second instinct was to actually think about it.
NCA -- Non-Compartmental Analysis -- is, at its core, a math problem. Trapezoidal integration, lambda-z regression, a handful of derived parameters. I've run these calculations hundreds of times by hand. I know what the outputs should look like. I know which edge cases will break a naive implementation: irregular sampling intervals, BLQ (Below Limit of Quantification) values, and flip-flop kinetics.
The math isn't proprietary. But the license fee sure is.
🔧 The Domain Wasn't the Hard Part
I'm a pharmacist. I already knew the pharmacokinetics. That part was fine.
The hard part was everything I didn't know about building software:
- Floating-point arithmetic misbehaves in ways that pen-and-paper PK never warned me about. Numbers that should be equal suddenly aren't. It's humbling.
- Edge case handling: what commercial software shows as a polite warning dialog needs to become explicit logic in code. Silent wrong answers aren't an option when someone's using this for bioequivalence work.
- Validation infrastructure: I couldn't just compare outputs to my hand calculations and call it done. I needed to match FDA-cited textbook references. The gold standard, not the "good enough" standard.
Knowing what "correct" looks like before writing code turned out to be a massive advantage. Most developers building scientific tools don't have that. Most scientists building tools don't have the engineering side. I had half of each, which is either the worst or the best starting position depending on the day.
🤖 Where AI Actually Helped (and Where It Didn't)
I need to be clear here, because "AI built my PK tool" isn't just wrong -- it's the kind of claim that makes regulatory people break out in hives. And they'd be right to.
What AI helped with:
- Translating math to code -- expressing terminal elimination regression in a way that's maintainable and readable
- Surfacing implementation pitfalls -- numerical issues in interpolation and rounding that I wouldn't have caught until something produced a wrong answer at 3 AM
- UI layout -- table structures that researchers can actually read without squinting or tilting their monitors
What AI can't do: tell me whether the output is correct. PK calculations have right and wrong answers. Knowing the difference is the job, and that's not something you can delegate to a language model.
✅ The Milestone That Mattered
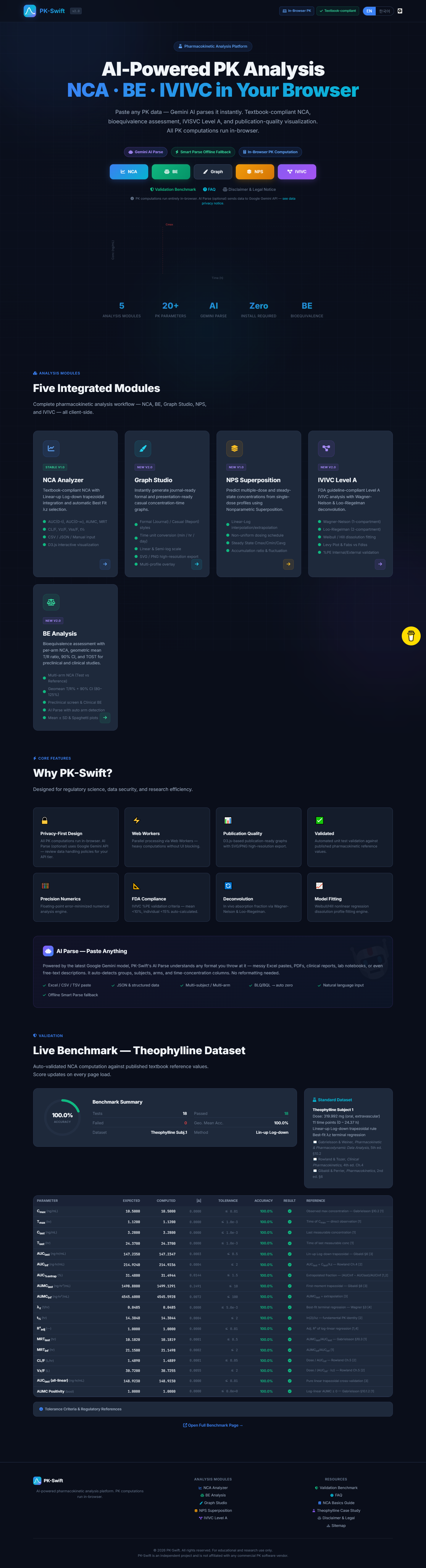
PK-Swift's AUC values now match FDA-cited textbook examples to 4 decimal places.
That's not me showing off. In bioequivalence work, precision isn't optional. A rounding difference in the fourth decimal can change whether a generic drug gets approved. The entire point of this project is that the math should be accessible to everyone -- not locked behind a $30K license.
I validated against the classic example from Gibaldi and Perrier's Pharmacokinetics, cross-checked my trapezoidal AUC against published bioequivalence datasets, and specifically tested the lambda-z (terminal elimination rate constant) calculation -- the parameter that trips up half of naive implementations because of how sensitive it is to the final few data points. When I got 0.4832 hr⁻¹ and the reference case showed the same, I knew the math was solid.
💡 What I'd Tell Someone Building Scientific Tools Without a CS Background
- Write acceptance criteria first. Define expected values and tolerated error ranges before you write a single line of code. "It looks about right" doesn't cut it in pharma.
- Build one golden dataset. Run it after every meaningful change. If the golden dataset breaks, you broke something. Simple.
- Keep assumptions visible. If BLQ is treated as zero, say it explicitly in the UI and the docs. Hidden assumptions are hidden bugs.
- Use AI as a sparring partner, not an authority. It's great at suggesting approaches. It's terrible at knowing when the approach is wrong for your domain.
- Version prompts and formulas in the same repo as code. Future-you will forget why decisions were made. Past-you should leave notes.
🚀 What's Next
Bioequivalence analysis: two-period crossover, geometric mean ratio CIs, and the 80-125% bounds.
The statistical layer is harder. The stakes are higher. And honestly, that's what makes it worth building carefully. If I rush this part, I'll end up with something that produces confident-looking numbers that are subtly wrong -- which is worse than producing no numbers at all. The NCA module is stable now -- I've spent the last month stress-testing against edge cases and squashing numerical precision bugs. Next is building the statistical inference layer with the same rigor.
2026.02.15
PK·SWIFT B.LOG - Follow along as each experiment unfolds.
Written by
Jay Lee
Korea-Licensed Pharmacist (#68652) · Senior Researcher
Korea University, College of Pharmacy (B.S. + M.S., drug delivery systems & industrial pharmacy). Building production-grade AI tools across medicine, finance, and productivity — without a CS degree. Domain expertise first, code second.
About the author →Related posts